These days, corn seems to be going high tech, as the U.S. and other countries shift their attention to corn-based ethanol fuels in response to dwindling oil supplies. And the corn market is feeling this demand for the “fuel of the future”: in recent months, corn prices have skyrocketed, in Mexico the cost of tortillas has soared, and many U.S. farmers have planned to invest more of their fields in corn, passing up soybeans, wheat, and cotton. So is corn the next new “green” technology? Well, maybe — but in fact, corn as we know it has always been high-tech. That plump ear of corn in the farmer’s market is just as much a product of human engineering as an iPod. Ten thousand years ago, there was no corn — there was just the weedy grass teosinte. Since then, humans have used evolution’s tool-kit to shape teosinte into the tall stalks of modern corn, joining in the other green revolution — the 6000 years of human history during which we transformed our lifestyles and environments by evolving all the major crops that we depend upon today: corn, rice, wheat, squash, millet, barley, banana, taro, tomato, potato, beans, and cotton. Now, scientists are learning more about exactly how we accomplished that feat.
Where's the evolution?
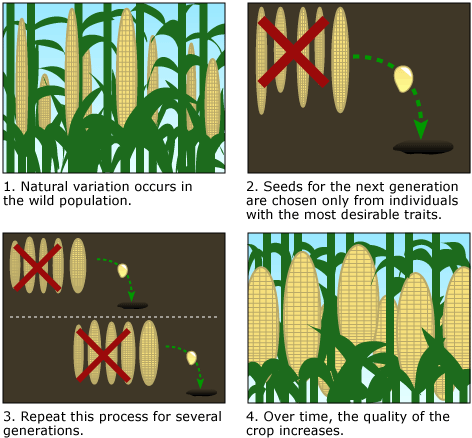
Plants and animals are domesticated through artificial selection, which works like natural selection does, but with humans instead of nature doing the selecting. As a simple example, imagine a stand of teosinte plants. The population varies: some plants are taller, some shorter, some have plumper seeds, some puny seeds, some have more seeds, some fewer (see step 1 below). Which would you prefer to eat next year? Perhaps, you collect some of the plump seeds and plant them in a convenient location (step 2 below). The plants that grow from those seeds carry more genes for plumpness (and have plumper kernels), but they still vary in many ways. That year, you collect seeds from the few plants in your crop with particularly plump kernels that are easiest to pick off and eat (step 3 below). The plants that grow from those seeds carry more genes for plumpness and for accessible kernels…and so on. Over many years of selection, the frequency of desirable gene variants increases in the population — and so does the quality of the crop (step 4 below).
This basic process has produced all of our major crop plants from their wild ancestors, some of which are shown here. Now scientists are trying to reconstruct important events in the domestication process. By figuring out how humans improved crops in the past, scientists may uncover valuable clues about how to improve crops even further to meet changing human needs. Most importantly, they would like to figure out which gene variants were favored by ancient people (i.e., which genes were the targets of selection). But this is a puzzle since we lack a blow-by-blow record of the long, slow process of domestication. In many cases, we are left with just the modern crop and its wild cousin; the two are obviously different, but how did we get from one to the other?



In December 2006, researchers John Doebley, Brandon Gaut, and Bruce Smith summarized much of what we currently know about the genes important in crop domestication and highlighted two basic approaches for identifying them: plant-down and gene-up. In the plant-down approach, researchers start with the phenotype — the physical features — of the plant and pick out traits that seem likely candidates for selection: rows of kernels per ear, number of ears per plant, etc. The researchers then look for associations between particular traits and particular sequences in the genome: if a trait and a sequence always seem to show up in the same plant, the gene sequence likely encodes that trait.
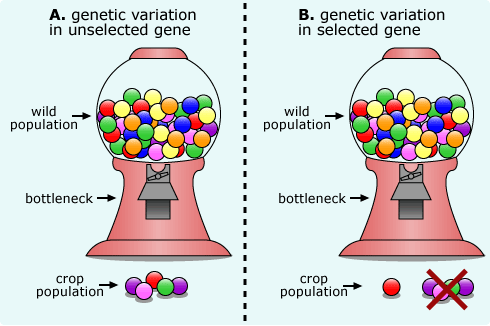
The gene-up approach, however, relies on an understanding of how artificial selection works on a population. Wild populations, like teosinte, have a lot of genetic variation — there are many different versions of many different genes in the population. When humans start to domesticate that plant, they begin with a small subset of the wild population. Since so few individuals are chosen as the founding fathers of the would-be crop plant, many of the gene versions present in the original wild population are not represented in the subset of the population beginning the process of domestication. It’s a bit like getting candy out of a gumball machine: the machine (i.e., the wild population) might have a great variety of gumball colors, but if you only get five pieces of gum (i.e., the starting population for the crop), you are unlikely to sample the full variety of gum colors. In genetics, this situation is called a bottleneck and it causes a large reduction in genetic variation for the incipient crop (as shown in scenario A below).
Because of this bottleneck, all the genes in the crop population, will tend to have less genetic variation that their counterparts in the wild population. However, the situation is even more extreme for the genes that are targeted for artificial selection. For these genes, ancient people only allowed plants carrying desirable gene versions to reproduce. This is a bit like going to the gumball machine when you only like cherry-flavored gum: you might reject many pieces of gum based on their color and will wind up with a handful of gumballs with very little variety. Genes that are the targets of selection lose even more genetic variation than others and are often reduced to the one gene version that produces the desirable trait (as shown in scenario B below).
The gene-up approach for identifying genes important in domestication is based on this difference in genetic variation: all genes in domestic crops have less genetic variation than their wild counterparts — but we can identify artificially-selected genes because they have unusually low genetic variation. To use this approach, a researcher needs a lot of data: many gene sequences from many different plants in the wild population and many gene sequences from many different plants in the domestic population. He or she then picks out the genes that meet this profile (unusually low plant-to-plant variation in the domestic population) and tries to figure out the function of those genes.
Either way, gene-up or plant-down, the point of all this work is to identify the genes that have played a role in the evolution of domestic crops from their wild ancestors. This may seem a bit esoteric — after all, why should we care which plants traits caught the fancy of prehistoric people? In fact, scientists care about this for very contemporary reasons. Understanding these genes and their other wild variants may hold the key to genetically engineering even more useful varieties of our crop plants. The more we learn about these important genes, the greater the possibilities for developing varieties that are more disease resistant, more nutritious, better tasting, more environmentally sound, or even — who knows? — honed to feed the growing ethanol market!
Primary literature:
- Doebley, J. F., Gaut, B. S., and Smith, B. D. (2006). The molecular genetics of crop domestication. Cell 127(7):1309-1321. Read it »
News articles:
- A kid-friendly tutorial on corn domestication from the Koshland Science Museum
- A summary of recent research on corn's ancestry from the National Science Foundation
- An article on one of the key genes that distinguish corn and teosinte from the University of Wisconsin-Madison
Understanding Evolution resources:
- Review the process of natural selection here. Explain the differences and similarities between the processes of natural selection and artificial selection.
- Look back at the photo of the wild and domestic tomatoes shown in this article. In a series of steps, explain how ancient people might have evolved a population of domestic tomatoes from a population of wild ones. Be sure that your explanation includes the concepts of variation, inheritance, and selection.
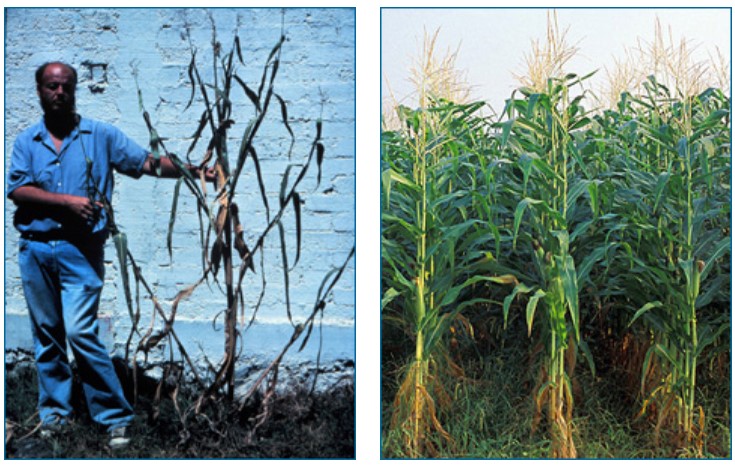
- Compare the pictures of wild teosinte and domesticated corn shown in this article. What trait variants in teosinte do you think would have been selected by ancient people? List at least three traits that you think might have been a target of selection and might have been involved in the process of domestication.
- What is a genetic bottleneck? Why does domestication result in a bottleneck? What else (besides domestication) might cause a bottleneck?;
- What is the effect of a genetic bottleneck on the genetic variation in a population?
- Imagine that you are researcher trying to use the “gene-up” approach to identify genes involved in domesticating tomatoes. You collect the following data:
Gene X variants Gene Y variants Gene Z variants Domestic tomatoes X2, X3, X5 Y4 Z1, Z3, Z5, Z6 Wild tomatoes X1, X2, X3, X4, X5 Y1, Y2, Y3, Y4, Y5, Y6, Y7 Z1, Z2, Z3, Z4, Z5, Z6 Which gene, X, Y, or Z, was most likely a target of selection in the process of domestication? How do you know?
- Teach about selection: In this second webcast of a four-part series appropriate for grades 9-12, evolutionary biologist David Kingsley discusses the processes of natural and artificial selection and how just a few small genetic changes can have a big effect on morphology, using examples from corn, dog breeding, and stickleback fish.
- Teach about bottlenecks: In this lesson for grades 9-12, students achieve an understanding of the Hardy-Weinberg Equilibrium without recourse to algebra by using decks of playing cards. An extension to the activity allows the teacher to simulate the effect of a bottleneck on a population.
- Teach about genetic variation: In this lesson for grades 9-12, students explore the natural variations present in a variety of organisms by examining sunflower seeds and Wisconsin Fast PlantsTM to consider the role of heredity in natural selection.
- Teach about corn domestication: This website provides educational resources related to the Doebley team's research. It includes a downloadable slide show appropriate for grades 6-12 on the major steps in the domestication of wild teosinte plants.
- Doebley, J. F., Gaut, B. S., and Smith, B. D. (2006). The molecular genetics of crop domestication. Cell 127(7):1309-1321.
- Ethanol's effect: Expensive tortillas. (2007, January 13). Chicago Tribune. Retrieved January 19, 2007 from Chicago Tribune.
- Seewer, J. (2007, January 19). Farmers planting more corn this year. The Washington Post. Retrieved January 19, 2007 from The Washington Post.