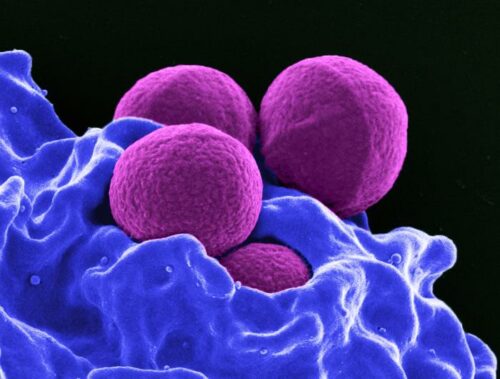
So-called superbugs, germs resistant to one or more of the medications meant to kill them, represent a new villain in the global fight for human health and welfare. Many of the “wonder” drugs discovered in the 1950s, which once seemed to promise the end of infectious disease, are no longer effective against many microbes. For example, Staphylococcus aureus, a common source of skin infections, used to be easily treated with penicillin, but today many strains are resistant to that and several other antibiotics, making a minor scratch potentially life-threatening. Drug-resistant bacteria, viruses, fungi, and parasites cause more than 700,000 deaths per year worldwide. In many cases, contracting a resistant strain of a pathogen more than doubles one’s chance of dying of the infection. Pharmaceutical companies are pivotal in combatting this global health emergency — and last month the Access to Medicine Foundation announced a new way to track drug companies’ efforts in this fight. Consumers, policymakers, and other drug companies can consult the Antimicrobial Resistance Benchmark to find out what companies are doing (and which are doing the most) to ensure that antimicrobials remain a useful part of our medical arsenal. But how exactly does a drug company fight superbugs?
Where's the evolution?
The first step in curbing resistant pathogens is recognizing that they are the result of a real-time evolutionary process. Because pathogens often have high mutation rates, a large population contains many different random genetic variants — so many, in fact, that one or more of them is likely to confer resistance to an antimicrobial drug. When the germs are exposed to medications (e.g., when a teen with a staph infection takes penicillin), most of the bugs will die or stop reproducing, but the few that carry gene versions that allow them to resist the antimicrobial will survive and pass the genes for resistance on to the next generation. Resistant strains evolve through this process of natural selection and, if the drug is widely used, become common over time. Because pathogenic life cycles are so short (an E. coli population can double in size every 17 minutes), this can occur quickly. In the case of the antibiotic streptomycin, evidence for the evolution of resistant tuberculosis bacteria was found in its first clinical trial. The challenge is made even more daunting by the fact that gene versions can be passed directly from one bacterium to another. Through this process of horizontal transfer, sets of genes for resistance to multiple antibiotics can be passed to a bacterial species that may not have been treated with any of the relevant drugs itself. It also allows resistance genes that evolved in, say, a pig farm where the animals are treated with antibiotics, to be transferred to another lineage and ultimately wind up in a bacterium that threatens human health. Fighting resistant microbes then means finding ways to slow the evolution of resistance in the first place and to prevent resistance genes from being shared too widely.
So what can drug companies do to combat microbial evolution and maintain an arsenal of useful, infection-fighting medications? The Antimicrobial Resistance Benchmark evaluates companies on a few different dimensions:
- Developing new antibiotics with new modes of action. This is important because evolution has no foresight: natural selection favors genes for resistance to antimicrobials that are in the environment right now. Resistance to our most commonly used antibiotics is already widespread, but a new antimicrobial that works in a different way from others is like a clock reset, starting with zero resistant strains. The Benchmark also gives credit for collaboration on drug development, since sharing ideas and information can lead to more creativity and insight, and speed up the rate at which new antimicrobials are available.
- Producing high-quality antimicrobial drugs. Antibiotics that are less effective can potentially speed the evolution of resistance. That’s because resistance genes come in a range of strengths and can be fine-tuned into a stronger form by evolution. While a high-quality antibiotic might wipe out virtually all the bacteria, a low-quality one could let a range of partially resistant individuals survive to reproduce. Each generation of reproduction is a chance for random mutation to introduce variants on the genes the individuals currently carry, including that gene for partial resistance. Over generations, this process can lead to a lineage that is strongly resistant, so that when a high-quality drug is encountered, it is ineffective. Companies get credit for being committed to good manufacturing practices.
- Ensuring that antimicrobials are used effectively, but only when needed. The more antimicrobials are used, the more strongly resistant strains are favored and the more quickly resistance evolves. It is particularly important that antimicrobials only be used on pathogens that are vulnerable to them (e.g., no antibacterial drugs for viral infections, no clindamycin for a clindamycin-resistant strain). Though you may take a drug to fight a specific pathogen, many other bacteria and viruses in your body will be exposed to that drug and experience selection favoring the evolution of resistant forms. And because many pathogens can share genes horizontally, a resistance gene that evolves in the harmless residents of your gut could easily make its way into a more malicious invader. The Benchmark encourages targeted, limited use by giving credit to drug companies that discourage their sales teams from focusing on volumes of antimicrobials sold, support doctor education on when different medications should be prescribed, label the medications clearly so that patients don’t overuse them, and develop plans for new drugs so that they are used appropriately as soon as they enter the market. Some companies are even restricting the use of new drugs to the highest priority infections. For example, Johnson & Johnson’s new antibiotic bedaquiline is distributed in a tightly monitored way and used only to treat tuberculosis infections that are already resistant to multiple other antibiotics. This approach, coupled with doctor education on the drug, will help ensure that bacterial populations are not needlessly exposed to the drug and to the natural selection that comes along with it.
- Keeping antibiotics out of waste. When factories release antimicrobials into waterways along with wastewater, the drugs can make their way into the groundwater and household taps, unnecessarily exposing many germs to this selective pressure. Even when the drugs do not directly reach humans, soil- and water-living bacteria are exposed to them and experience natural selection favoring the evolution of resistance genes. Though these microbes do not themselves threaten human health, the resistance genes can be picked up by pathogenic bacteria via horizontal transfer and render the drug that the factory produces less useful. Companies get credit for limiting and monitoring antibiotics that are released in wastewater during drug production.
- Monitoring pathogen evolution. In order to fight the evolution of resistant microbes, we need to know where this is occurring and how quickly. The Benchmark gives credit to companies that participate in and support resistance surveillance programs.
When all of these factors are taken into account, the big winner among large, research-based drug companies is GlaxoSmithKline, which racked up more than 75% of the points available for doing its part to slow the evolution of resistant pathogens. The worst performer, Roche, scored less than 50%.
The Antimicrobial Resistance Benchmark is a way to recognize and share the practices of pharmaceutical companies’ work to preserve and expand the utility of our infection-fighting drugs, and this tally provides companies with a starting point to improve their policies. But fighting the evolution superbugs is a big job, and evolution doesn’t have a pause button. New medications will be vulnerable to the rise of resistant pathogens, just as our old ones were. Hence, keeping ahead in this ongoing struggle will require many different stakeholders working together — doctors, patients, consumers, farmers, policymakers, and drug companies. From the pharma executive who decouples employees’ bonuses from volumes of vancomycin sold, to the supermarket shopper who chooses to buy pork raised without antibiotics, we all have a role to play.
Primary literature:
- Access to Medicine Foundation. (January, 2018). Antimicrobial Resistance Benchmark 2018. Amsterdam, Netherlands. Read it »
News articles:
- A report on the new index from the New York Times
- A summary of the Benchmark report from Science Magazine
Understanding Evolution resources:
- Explain how mutation and horizontal transfer affect the genetic variation in a microbe population.
- Review the process of natural selection. Explain why genetic variation is so important to this process.
- The components of natural selection can be summarized as: variation, selection, inheritance, and time.
- Which of the five dimensions captured by the Antimicrobial Resistance Benchmark focus on strengthening the action of selection on pathogens? Explain your answer.
- Which focus on reducing pathogens’ exposure to selection for resistance? Explain your answer.
- Come up with a list of at least three things that an individual consumer or patient can do to slow the evolution of drug resistant pathogens.
- Explain how evaluating companies on their efforts against superbugs could impact the evolution of drug resistance.
- Teach about natural selection and antibiotic resistance: In this activity for grades 9-12, students learn why evolution is at the heart of a world health threat by investigating the increasing problem of antibiotic resistance in such menacing diseases as tuberculosis.
- Teach about the genetic basis of antibiotic resistance: In this case study for college students, learners compare the genomes of MRSA and its genetic cousin MSSA to locate DNA differences and mobile elements that could be expected to improve the bacterium's resistance to antibiotics and its ability to cause disease.
- Access to Medicine Foundation. (January, 2018). Antimicrobial Resistance Benchmark 2018. Amsterdam, Netherlands. Retrieved March 5, 2018 from https://amrbenchmark.org/wp-content/uploads/2018/01/Antimicrobial-Resistance-Benchmark-2018.pdf
- D’Arcy Hart, P. (1999). A change in scientific approach: from alternation to randomised allocation in clinical trials in the 1940s. BMJ. 319: 572-573.
- Davies, J. (2006). Where have all the antibiotics gone? Canadian Journal of Infectious Diseases and Medical Microbiology. 17: 287-290.
- McNeil Jr, D. G. (January 23, 2018). New index rates drug companies in fight against 'Superbugs.' The New York Times. Retrieved March 5, 2018 from https://www.nytimes.com/2018/01/23/health/antibiotic-resistence-glaxo-johnson.html
