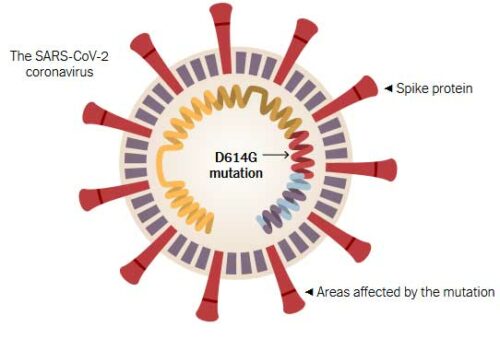
Early in the pandemic, a particular mutation in the virus that causes COVID-19 (SARS-CoV-2) caught scientists’ attention. The mutation, known as D614G, makes a small change to the virus, causing a single amino acid in the spike proteins of the virus to be swapped for a different amino acid. After the mutation arose (likely in China), it spread, and lineages carrying this mutation came to dominate many outbreaks. This left scientists wondering if the D614G mutation is an adaptation that helped the virus along as it spread around the world. Since then, every month brings a new slew of studies on the mutation, along with alarming headlines. September 2020 was no exception. The Washington Post declared about a recent D614G investigation, “Massive genetic study shows coronavirus mutating and potentially evolving amid rapid U.S. spread.” And yet, many scientists question whether D614G is anything notable. Here, we’ll briefly explain the lines of evidence and a few different evolutionary explanations for what’s going on.
Where's the evolution?
First, there is no question about whether or not the new coronavirus is evolving as it spreads through human populations. It is. New mutations are arising in the virus all the time. Most of those are detrimental to the virus and are immediately weeded out. Furthermore, different viral strains with different genetic sequences are becoming more or less common just by chance. That alone is evolution. All populations evolve over time via mutation and genetic drift. There’s nothing inherently scary about that. The real question here is whether D614G has reached its high frequency through natural selection.
Mutation and genetic drift don’t reliably shape the evolution of a population’s traits in any particular way. If mutation and genetic drift are solely responsible for the virus’ recent evolution, then the virus likely has about the same effect on humans today as it did back in January. But if the virus is also evolving through natural selection, it could shape the pandemic in worrisome ways.
Natural selection spreads traits that enable an individual to reproduce more — that is, to get more copies of their genes into the next generation. What sort of traits is natural selection likely to favor in a pathogen? It is sometimes suggested that pathogens will evolve into less deadly (i.e., less virulent) forms over time because a host that dies quickly won’t pass the disease on, thwarting its reproduction. But clearly, that depends on the particulars of the infection and the people infected. Cholera, for example, spreads through water, and doesn’t require one live person to come into contact with another. And Ebola can be contracted even from a dead body. In terms of COVID-19, there is no strong evidence that the D614G mutation makes the coronavirus more or less deadly (and scientists are certainly looking for that evidence if it’s out there!). However, some evidence hints that D614G could help the virus spread more easily, and that has been the topic of much scientific debate.
So what’s the evidence? There are a few relevant lines, and each comes with its own set of caveats:
- Viral and protein structure. The D614G mutation affects a spike protein. The virus uses its spikes to enter cells. So it seems plausible for a change like D614G to affect transmission. Of course, there are probably plenty of changes to the spike protein that would have no effect on transmission. But, at the very least, a mutation that affects the spike protein seems more likely to boost transmission than a mutation that affects, say, a protein deep inside the virus particle with no obvious role in infecting new cells.
- Studying virus transmission among cells. Cells infected with D614G viruses passed the virus on to more other cells, in comparison to cells infected with a viral strain that did not carry the mutation. So far, because of the safety precautions needed to study the coronavirus itself, these tests have used other viruses engineered to display SARS-CoV-2’s cell-entry spike. This finding is suggestive — but at the same time, it’s a big leap from non-coronavirus cells in a figurative petri dish to humans infected with COVID-19 out in the world.
- Observing viral loads in patients. One recent study looked back at testing data from patients diagnosed with COVID-19 and found that, at the time the test was done, patients carrying the D614G virus had more copies of the virus in their nose/throat area than did patients who had a strain of the virus without the mutation. This evidence suggests a mechanism for a faster rate of spread. Perhaps patients with a higher viral load in their noses and throats release more virus into the air when they breathe, talk, and cough, and so infect more people. But again, this data doesn’t actually show that this happens; it just suggests it’s possible.
- Patterns of gene frequency over time. Many studies have examined how different viral strains fare in outbreaks of COVID-19. Does the D614G strain consistently spread to more people than do strains without the mutation? Several studies have found that it does. That might sound like an open and shut case: of course, the mutation makes the virus more transmissible. But in fact, this pattern might also be observed if the mutation has no effect on the virus’ ability to spread. Here’s what’s going on. When a small population expands into a new habitat (or, in this case, when SARS-CoV-2 starts spreading in a new community), the population size (i.e., of the virus) increases rapidly. And in the process, it’s very likely that just one or a few members of the initial population (or, in this case, early SARS-CoV-2 infections) will leave behind many more descendants than others, just by chance. This is a form of genetic drift called the founder effect. The more common one gene version or mutation is in the founding population, the more likely it will end up being prevalent later on. The D614G mutation arose early in the pandemic and was carried out of China in the first waves of spread. It had a big advantage over other strains based on that alone. Is its dominance now because it is actually better at spreading or just because it was in the right places at the right times? Figuring that out is challenging. Some of the studies that try to seem to have found a genuine advantage to D614G, but others have not.
In all of these cases, scientists are evaluating both sides of evidence, and so they have not yet reached a consensus. Making things even more difficult to sort out, most of the findings described above have come from preprints — reports on research that have not yet been peer-reviewed and may not be in their final form. Once experts in each field have had a chance to weigh in and offer feedback, it might change how we interpret the evidence.
Even though scientists continue to debate whether or not D614G has spread through natural selection, there’s no question that these lines of research are worthwhile. Studies of spike structure and how the virus invades cells build knowledge that can help us develop new treatments and vaccines. And studies of person-to-person transmission and outbreaks can help us develop policies and interventions to slow the spread of the disease. And finally, viruses carrying D614G are now by far the most common strains of SARS-CoV-2, no matter how it got to be that way; fighting COVID-19 today means fighting viruses that nearly all carry this mutation.
Primary literature:
- Long, S. W., Olsen, R. J., Christensen, P. A., Bernard, D. W., Davis, J. J. Shukla, M., ... and Musser, J. September 29, 2020. Molecular architecture of early dissemination and massive second wave of the SARS-CoV-2 virus in a major metropolitan area. Preprint retrieved from medRxiv, September 29, 2020. Read it »
- Volz, E. M., Hill, V., McCrone, J. T., Price, A., Jorgensen, D., O'Toole, A., ... and The COVID-19 Genomics UK Consortium. September 1, 2020. Evaluating the effects of SARS-CoV-2 spike mutation D614G on transmissibility and pathogenicity. Preprint retrieved from medRxiv, September 29, 2020. Read it »
News articles:
- An article about a recent study of the mutation’s spread, from the Washington Post
- An explanation of why scientists are still debating the cause of the mutation’s spread, from LiveScience
Understanding Evolution resources:
- Since the beginning of the pandemic, the D614G mutation has become more common. Does this mean that coronavirus has evolved over the last nine months? Explain your answer.
- Scientists are currently investigating whether the COVID-19 virus is evolving through natural selection. What viral trait(s) are they focusing on in these studies?
- What sort of traits are likely to spread via natural selection? What sort of traits are likely to spread via genetic drift?
- What are two possible explanations for the rise of the D614G mutation over time? Explain in your own words.
- Advanced: In what situations might you expect a virus to evolve increased virulence as it spreads through a population? In what situations might you expect a virus to evolve decreased virulence as it spreads? Explain your answers.
- Teach about random mutations and natural selection: In this activity for grades 9-12, students build and evolve and modify paper-and-straw "birds" to simulate natural selection acting on random mutations.
- Teach about how scientists study natural selection: In this advanced 4-lesson curriculum unit for the high school and college levels, students examine evidence to compare four different explanations for why many malarial parasites are resistant to antimalarial drugs; investigate how scientific arguments using G6PD data show support for natural selection in humans; design an investigation using a simulation based on the Hardy-Weinberg principle to explore mechanisms of evolution; and apply their understanding to other alleles that have evolved in response to malaria.
- Teach about genetic drift: This set of five PowerPoint slides for college courses features questions for problem-based discussion (i.e., open-ended questions that engage students with each other and with course material) and can be easily incorporated into lectures on genetic drift.
- Callaway, E. September 8, 2020. The coronavirus is mutating — does it matter? Nature. Retrieved September 29, 2020 from https://www.nature.com/articles/d41586-020-02544-6
- Long, S. W., Olsen, R. J., Christensen, P. A., Bernard, D. W., Davis, J. J. Shukla, M., ... and Musser, J. September 29, 2020. Molecular architecture of early dissemination and massive second wave of the SARS-CoV-2 virus in a major metropolitan area. Preprint retrieved from medRxiv, September 29, 2020. DOI: https://doi.org/10.1101/2020.09.22.20199125
- Mooney, C., Achenbach, J., and Fox, J. September 23, 2020. Massive genetic study shows coronavirus mutating and potentially evolving amid rapid U.S. spread. The Washington Post. Retrieved September 29, 2020 from https://www.washingtonpost.com/health/2020/09/23/houston-coronavirus-mutations/?arc404=true
- van Dorp, L. September 24, 2020. Coronavirus mutations: what we've learned so far. LiveScience. Retrieved September 29, 2020 from https://www.livescience.com/coronavirus-mutations-explained.html
- van Dorp, L., Richard, D., Tan, C. CS., Shaw, L. P., Acman, M., Balloux, F. August 19, 2020. No evidence for increased transmissibility from recurrent mutations in SARS CoV-2. Preprint retrieved from bioRxiv, September 29, 2020. DOI: https://www.biorxiv.org/content/10.1101/2020.05.21.108506v5
- Volz, E. M., Hill, V., McCrone, J. T., Price, A., Jorgensen, D., O'Toole, A., ... and The COVID-19 Genomics UK Consortium. September 1, 2020. Evaluating the effects of SARS-CoV-2 spike mutation D614G on transmissibility and pathogenicity. Preprint retrieved from medRxiv, September 29, 2020. DOI: https://doi.org/10.1101/2020.07.31.20166082
- Yukovetskiy, L., Wang, X., Pascal, K. E., Tomkins-Tinch, C., Nyalile, T., Wang, Y., ... and Luban, J. July 16, 2020. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Preprint retrieved from bioRxiv, September 29, 2020. DOI: https://doi.org/10.1101/2020.07.04.187757
